Bio
The turtle module provides a visual and intuitive way to simulate biological structures like atoms, molecules and proteins.
Molecular Geometry
The Water Molecule (H2O)
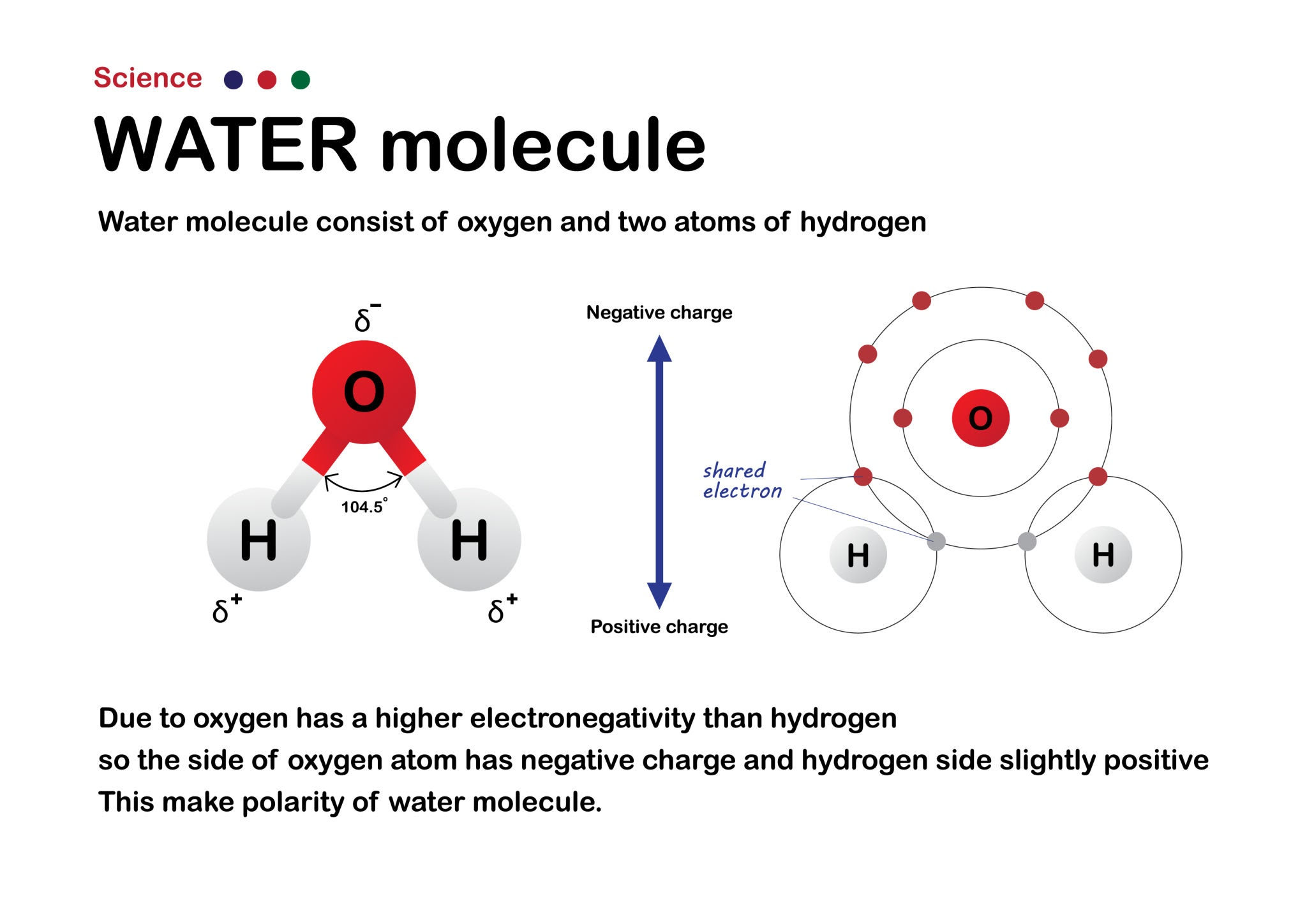
Draw a classic Mickey Mouse-shaped water molecule.
- Create a turtle to represent the drawing pen.
- Draw a large red circle to represent the Oxygen atom.
- Lift the pen, move to the center of the Oxygen atom, and set the turtle’s heading.
- Draw a thick gray line (the chemical bond) outward, then draw a smaller white (or light gray) circle for the first Hydrogen atom.
- Return to the center of the Oxygen atom. Turn the turtle exactly 104.5 degrees (the bond angle of water).
- Draw the second bond and the second Hydrogen atom.
Possible solution, draw Water Molecule (H2O):
# Simple Mickey Mouse Water Molecule (H2O)
=
# -------------------------
# Function to draw a filled circle (atom)
# -------------------------
# -------------------------
# Function to draw a bond + hydrogen
# -------------------------
# -------------------------
# Draw Oxygen
# -------------------------
# -------------------------
# Draw Hydrogens
# -------------------------
Biological Dynamics
Brownian Motion (Random Walk)
Simulate the erratic, random movement of a microscopic particle suspended in a fluid.
- Import the
randommodule alongsideturtle. - Create a small, colored dot (the turtle) representing a protein or molecule.
- Write a
whileloop that repeats 500 times. - Inside the loop, have the turtle turn a random angle (between 0 and 360 degrees) and move forward a random small distance (e.g., between 1 and 10 pixels).
- Leave the turtle’s pen down so it draws a trail of its path.
- Add an “invisible boundary” (like a cell membrane). If the turtle’s x or y coordinates go beyond a certain limit, force it to bounce back into the center.
Cellular Viral Infection Dynamics
Simulate a simple viral infection spreading through a cluster of cells.
- Create a list of 20 turtles. Scatter them randomly across the screen and stamp them as green circles. These are healthy cells.
- Create one red turtle (the virus/infected cell) and place it in the center.
- Create an animation loop where all turtles move randomly (using a mini random walk).
- The Dynamic Rule: Check the distance between the red turtle and the green turtles. If a green turtle gets within a specific distance (collision radius) of a red turtle, change the green turtle’s color to red.
- Watch as the “infection” dynamically spreads through the moving population!
You will need to use
turtle.distance()to calculate the proximity between the agents on the screen.
Possible solution
=
# --- Create green cells ---
=
=
# --- Create red cell (the virus) on center ---
=
# --- Custom params ---
= 20
= 10
# --- Main program ---
# Move green cells randomly
=
=
# Check infection
Protein Structures
Primary Structure (The Polypeptide Chain)
Draw a “beads on a string” model of a polypeptide chain using different colors to represent different types of amino acids (e.g., hydrophobic vs. hydrophilic).
- Create a list of colors in Python (e.g.,
["red", "blue", "green", "blue", "yellow", "red"]). Each color represents a specific amino acid. - Write a
forloop that iterates through your list of colors. - For each color, change the turtle’s fill color, draw a filled circle (the amino acid bead), and then move the turtle forward.
- Keep the pen down as you move forward to draw a black line connecting the beads—this represents the peptide bond.
First solution:
=
=
# --------------------------
# Sample AA (AminoAcids)
# hydrophobic = blue, hydrophilic = red, special = green
# --------------------------
=
# --------------------------
# Draw polypeptide chain.
# --------------------------
# començar a l'esquerra
# bead size
# go to next bead
Improved version, based on Real AA sequence. B chain of Human Hemoglobin. 17 AA.
# --------------------------
# Classe per a un aminoàcid
# --------------------------
=
=
=
# --------------------------
# Real AA sequence. B chain: Human Hemoglobin. 17 beads.
# --------------------------
# Format: (name, color)
= ,
,
,
,
,
,
,
,
,
,
,
,
,
,
,
,
]
=
=
# --------------------------
# Draw AA chain
# --------------------------
# Draw bead
# Save text position
=
# Draw lines.
# t.penup()
# t.pendown()
# --------------------------
# Write aminoacid name.
# --------------------------
, =
Secondary Structure (Alpha Helix & Beta Sheet)
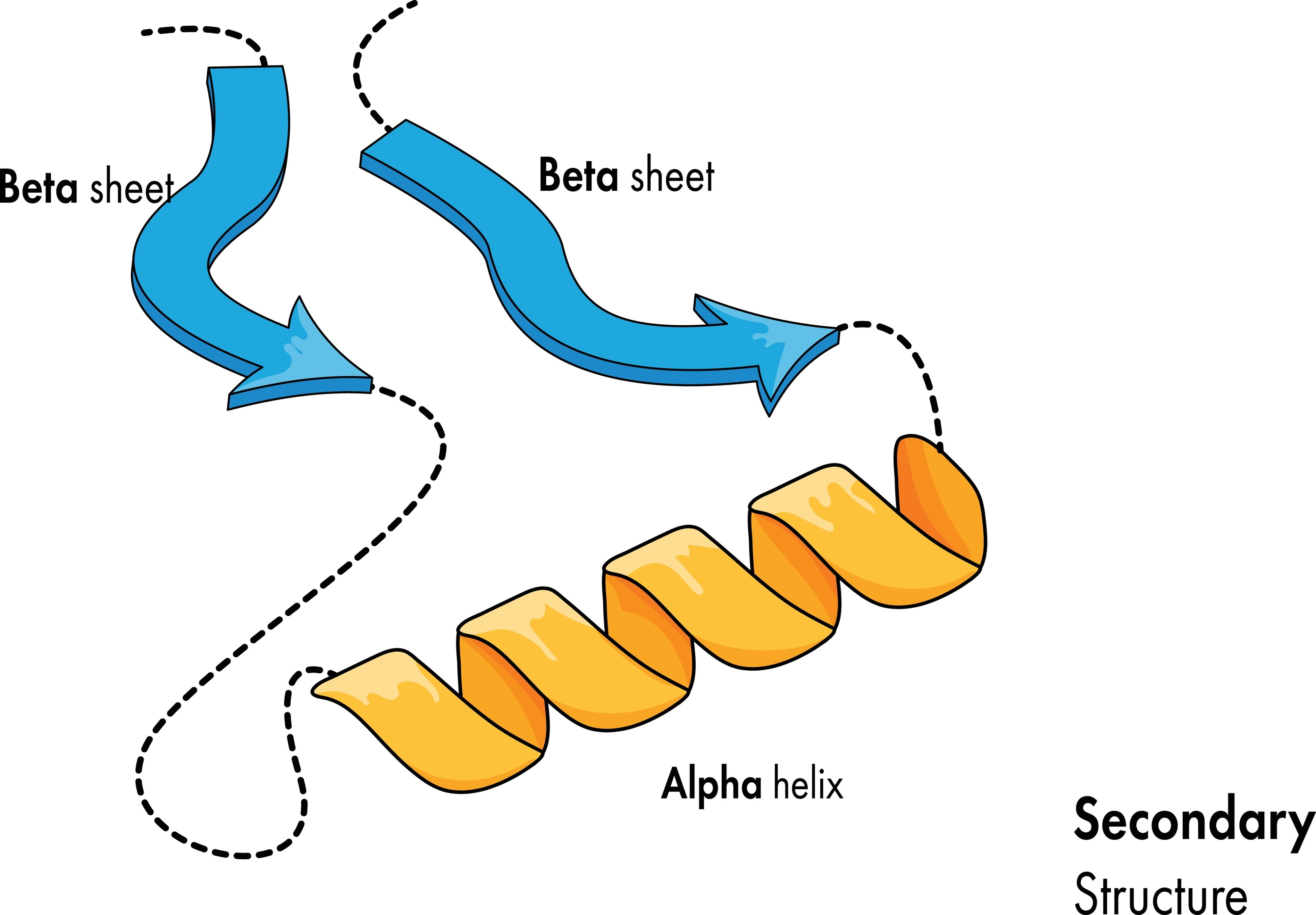
Write two separate functions to draw the two most common secondary structures side-by-side: a coiled alpha-helix and a zig-zagging beta-pleated sheet.
- The Alpha Helix: Write a loop that moves the turtle slightly forward and then draws a partial circle (using turtle.circle(radius, extent)). Repeating this slightly overlapping pattern creates a 2D illusion of a 3D spring or coil.
- The Beta Sheet: Lift the pen and move to a new area of the screen. Write a loop that uses sharp, alternating angles (e.g., turn left 60 degrees, move forward, turn right 120 degrees, move forward, turn left 60 degrees). Draw multiple parallel zig-zag lines to represent the “pleated” sheet.
- Draw small, dashed lines connecting the parallel strands of your beta sheet to represent the hydrogen bonds holding them together.
Repte: Dibuixar els aminoàcids i cadenes d’una proteïna d’un fitxer PDB.
Com recordaràs a https://xtec.dev/bio/protein/pdb podem descarregar-nos fitxers de proteïnes del Protein Data Bank i netejar-los
eliminant les linies que no ens interessen, i quedant-nos únicament amb les que defineixen la seva estructura.
Una de les possibles formes de descarregar-se una proteïna i deixar-la neta (sense aigues i altres residus) és aquesta:
import httpx
pdb_id = "1A4W"
url = f"https://files.rcsb.org/download/{pdb_id}.pdb"
response = httpx.get(url)
response.raise_for_status()
with open(f"{pdb_id}.pdb", "w") as f:
f.write(response.text)
print("PDB descarregat.")
with open(f"{pdb_id}.pdb") as infile, open(f"{pdb_id}_clean.pdb", "w") as outfile:
for line in infile:
if line.startswith("ATOM"): # només àtoms de la proteïna
outfile.write(line)
print("Proteïna neta (sense aigües ni HETATM).")També pots usar la eina Prody (que encara no hem vist amb profunditat):
- https://xtec.dev/bio/protein/structural-analysis
Llavors, a aquest codi cal afegir la forma de seleccionar la informació que ens interessa per a dibuixar-la amb Turtle.
El resultat esperat ha de ser semblant a aquest:
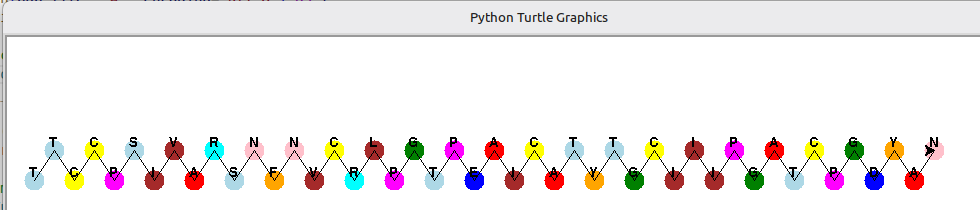
En el projecte de gitlab associat, dins del directori pdbeads, podràs analitzar una versió inicial.
Advanced Biological Dynamics
Enzyme-Substrate Complex (The Lock and Key)
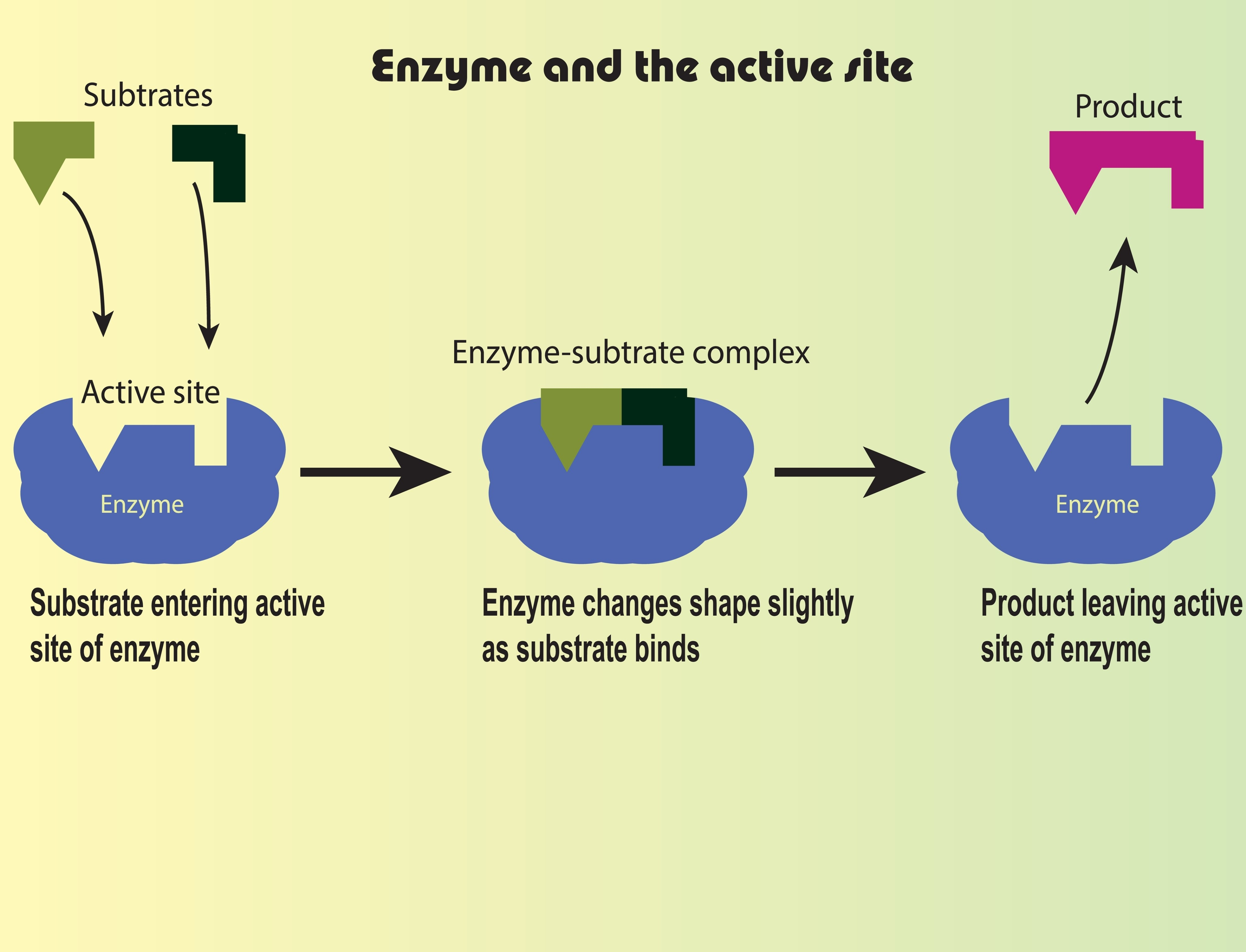
Animate a small molecule (the substrate) randomly floating around until it successfully “docks” into the specific shape of a larger molecule (the enzyme).
- Create a large, stationary turtle and use
turtle.shape()to give it a custom shape, or draw a “Pac-Man” like figure with an open mouth. This open mouth is the active site. - Create a smaller turtle (the substrate) that perfectly fits into that active site.
- Set the smaller turtle on a random walk loop (like in Exercise 3).
- The Dynamic Rule: Inside the loop, check the substrate’s coordinates. If it enters the exact coordinates of the enzyme’s active site, stop the random walk.
- Snap the substrate into the center of the active site, pause for 1 second (using
time.sleep()), change the substrate’s color (showing it has been catalyzed into a “product”), and have it bounce away.
Osmosis across a Semi-Permeable Membrane
Simulate water molecules moving to equalize concentration, while larger solute molecules remain trapped on one side.
- Draw a dashed vertical line down the exact center of the screen. This is your semi-permeable cell membrane.
- Create two lists of turtles. Make 30 small blue dots (water) and 10 large green dots (sugar/solute).
- Start the simulation with 15 water dots on the left side, and 15 water dots plus all 10 green dots on the right side.
- Animate all dots with a random walk.
- The Dynamic Rule: When a blue dot hits the dashed line’s x-coordinate, let it pass through. When a green dot hits the dashed line, force it to reverse its heading (bounce off), keeping it trapped on the right side.
- Observe over time: Because water moves freely, more water will eventually accumulate on the right side, simulating osmotic pressure!
To manage multiple moving turtles simultaneously without the animation lagging, turn off the automatic screen updates using
turtle.tracer(0)and manually update the screen at the end of each loop iteration usingturtle.update().
Cellular behavior
Chemotaxis is the movement of a cell in response to a chemical stimulus, and Phagocytosis is the act of a cell engulfing a large particle (like a white blood cell eating a bacterium).
The Macrophage Chase
Program a large white blood cell (the macrophage) to actively hunt down a small, erratically moving bacterium.
- Create a small, red turtle representing the Bacterium. Set it on a continuous, erratic random walk loop (moving fast, changing directions often).
- Create a large, white turtle representing the Macrophage.
- The Dynamic Tracking Rule: Inside the animation loop, have the Macrophage constantly check the
(x, y)coordinates of the Bacterium. - Use
macrophage.setheading(macrophage.towards(bacterium))so the white blood cell always points toward its prey. - Have the Macrophage move forward at a slightly slower speed than the Bacterium. Because the Bacterium moves randomly and the Macrophage moves directly, the Macrophage will eventually catch it.
- The Phagocytosis Rule: When
macrophage.distance(bacterium) < 20, hide the Bacterium turtle (usingbacterium.hideturtle()) and change the Macrophage’s color to indicate it has “digested” the pathogen.